 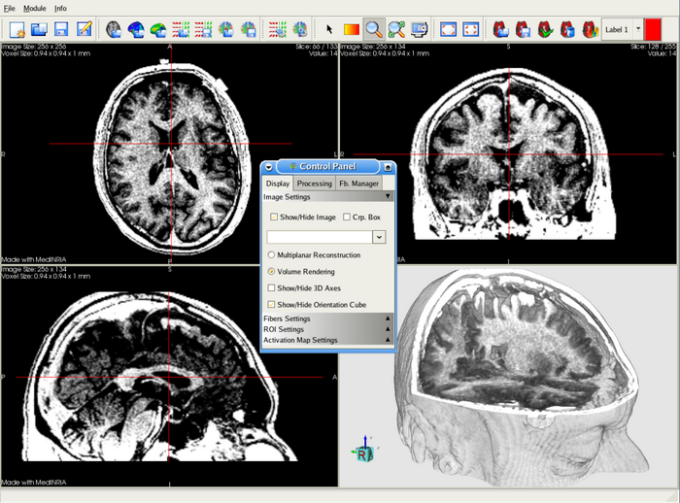
One b0 (i.e., b≈0) and 6 diffusion-weighted images (b≈1.5×10 3 s/mm 2) sensitized in 6 directions (,, ,, , ) were acquired over approximately 28 hours. Diffusion encoding was performed using a pair of half-sine gradient pulses (pulse width=1.3 ms, pulse separation=6.4 ms, gradient amplitude=1600 mT/m). A diffusion-weighted spin-echo pulse sequence was used to acquire 3D volume images (field of view=22×11×11 mm, matrix size=512×256×256 resulting in 43 μm isotropic resolution, TR=100 ms, TE=11.8 ms, NEX=2). Specimens were imaged in solenoid radiofrequency coil that was built in-house, constructed from a single sheet of microwave substrate. An additional stage in the transceiver chain converted the transmit and receive signals via a mixer driven with an external 400-MHz oscillator. Billerica, MA), controlled by a GE EXCITE MR imaging console (GE Healthcare, Milwaukee, WI). 
Īll images were acquired using a 9.4 T vertical bore Oxford magnet with shielded coils providing gradients of up to 2000 mT/m (Resonance Research, Inc. All of the data in this study is available at. The DTI data is now part of the Biomedical Informatics Research Network (BIRN) network ( Johnson et al., 2007), which enables investigators from any place on the globe to access quantitative mouse brain anatomy in a routine and standardized fashion. The coordinate system, called “Waxholm Space” (WHS) in reference to the Swedish city where this committee first met, has been developed to allow scientists to share many different types of data in a common reference space. This standardized coordinate system has been developed by the Digital Atlasing Task Force of the International Neuroinformatics Coordinating Facility. The DTI dataset of each individual brain is mapped to a common reference space defined by a collection of very high-resolution (21.5 μm) MR images with matching Nissl sections. The spatial resolution (80 pl voxel) allowed DTI parameters to be quantitatively measured at approximately 7 million locations throughout the mouse brain. 30 for the diffusion-weighted image (DWI) within 28 hours scan time.50 for the non-diffusion-weighted (b0) image, and SNR.In this study, streamlined methods for specimen fixation and staining, image acquisition, reconstruction, post-processing, and distribution have been devised to form an efficient protocol to obtain DTI datasets at 43 μm isotropic resolution with SNR Two technical issues make microscopic DTI challenging-the signal from the small voxels is very weak and the data arrays are exceptionally large. smaller voxels to reduce partial volume effects, resolve finer structures, and improve fiber tracking accuracy. Since DTI models all the tissues inside one voxel as a single ellipsoid, it is of great benefit to acquire higher-resolution data, i.e. Current state-of-the-art DTI methods enable imaging an ex vivo adult mouse brain at approximately 120 μm isotropic resolution (1700 pl voxel) within approximately one day of scan time ( Verma et al., 2005 Zhang et al., 2005). The scan time/SNR constraint is more prominent for high-resolution DTI, which is essential to image a small specimen such as the mouse brain. DTI provides a unique composite contrast reflecting tissue diffusivity, anisotropy, and directionality, which can also be used to track fibers ( Mori et al., 1999) to yield detailed three-dimensional connectivity information, offering a complementary alternative to the traditional histological methods.ĭue to the nature of signal attenuation in diffusion-weighted imaging and the requirement of multiple images with different diffusion weighting gradients, DTI requires a much longer scan time than conventional magnetic resonance imaging (MRI) to acquire a complete dataset with sufficient signal-to-noise-ratio (SNR). Diffusion tensor imaging (DTI) ( Basser et al., 1994) of the mouse brain has resulted in extraordinary new insight into the brain structure, connectivity, and integrity in normal and diseased brains, especially in white matter regions ( Mori et al., 2001 Song et al., 2005). Advances in molecular biology have provided unprecedented opportunities to use versatile small animal platforms (most importantly, the mouse) for connecting genotypes and phenotypes of brain development, damage, or repair associated with specific pathologies ( Badea et al., 2009 Chesler et al., 2003). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
